BioMicroCenter:SpTx: Difference between revisions
Resize images: workflow 250px, Boyer cardiomyocytes 300px |
Add Singular G4X in situ spatial sequencing section (beta stub) |
||
| (3 intermediate revisions by 2 users not shown) | |||
| Line 4: | Line 4: | ||
== 10X VISIUM HD == | == 10X VISIUM HD == | ||
[[File:VisiumHD_workflow_slide.jpg | {| width="100%" | ||
|- | |||
|style="width: 200px;"| | |||
[[File:VisiumHD_workflow_slide.jpg|left|250px|Workflow and capture areas for Visium HD]] | |||
<div style="clear:both; padding-top:4px; font-size:12px; color:#555;">Workflow and capture areas for Visium HD</div> | |||
| | |||
[https://www.10xgenomics.com/products/visium-hd-spatial-gene-expression 10x Visium HD] is a spatial transcriptomics platform that profiles whole-transcriptome gene expression across intact tissue sections at single-cell-scale resolution. Tissue sections are processed through the CytAssist instrument, which transfers RNA complexes onto a capture slide containing a continuous array of 3,000,000 spatially barcoded 2 µm squares covering a 6.5 × 6.5 mm capture area (~0.65 cm²). Two chemistries are available: a probe-based assay for human and mouse (FF, FxF, and FFPE), and a polyA-based assay (Visium HD 3') for fresh frozen tissue from any species. H&E or immunofluorescence (IF) staining can be performed prior to processing for protein co-detection. | [https://www.10xgenomics.com/products/visium-hd-spatial-gene-expression 10x Visium HD] is a spatial transcriptomics platform that profiles whole-transcriptome gene expression across intact tissue sections at single-cell-scale resolution. Tissue sections are processed through the CytAssist instrument, which transfers RNA complexes onto a capture slide containing a continuous array of 3,000,000 spatially barcoded 2 µm squares covering a 6.5 × 6.5 mm capture area (~0.65 cm²). Two chemistries are available: a probe-based assay for human and mouse (FF, FxF, and FFPE), and a polyA-based assay (Visium HD 3') for fresh frozen tissue from any species. H&E or immunofluorescence (IF) staining can be performed prior to processing for protein co-detection. | ||
<BR><BR> | <BR><BR> | ||
|} | |||
{| | {| | ||
| Line 85: | Line 90: | ||
=== 10X Visium HD === | === 10X Visium HD === | ||
[[File:VisiumHD_loupe_example.jpg|thumb|right| | [[File:VisiumHD_loupe_example.jpg|thumb|right|350px|Example of FF data from Visium HD]] | ||
'''FAQs for Users'''<BR> | '''FAQs for Users'''<BR> | ||
| Line 117: | Line 122: | ||
|} | |} | ||
== AVITI24 CYTOPROFILING & IN SITU SEQUENCING == | == AVITI24 CYTOPROFILING & IN SITU SEQUENCING == | ||
<div style="background:#fff8e6; border-left:4px solid #c8960c; padding:8px 12px; margin-bottom:12px;">'''Beta Service — Early Access:''' The AVITI24 Teton CytoProfiling and DISS services at the BioMicro Center are operational but not yet routine | <div style="background:#fff8e6; border-left:4px solid #c8960c; padding:8px 12px; margin-bottom:12px;">'''Beta Service — Early Access:''' The AVITI24 Teton CytoProfiling and DISS services at the BioMicro Center are operational but not yet routine. '''Consultation with BMC staff is required before submitting samples.'''</div> | ||
<!-- placeholder image left: AVITI24 instrument photo or Teton CytoProfiling schematic --> | <!-- placeholder image left: AVITI24 instrument photo or Teton CytoProfiling schematic --> | ||
<!-- replace with: [[File:AVITI24_instrument.png|thumb|left|400px]] --> | <!-- replace with: [[File:AVITI24_instrument.png|thumb|left|400px]] --> | ||
| Line 222: | Line 227: | ||
<BR><BR> | <BR><BR> | ||
'''Pricing'''<BR> | '''Pricing'''<BR> | ||
See [[BioMicroCenter:Pricing|BioMicroCenter:Pricing]] → Single Cell & Spatial Services → AVITI24 for current pricing. Sequencing-only runs on the AVITI24 are charged at standard AVITI24 sequencing rates. | |||
<BR><BR> | <BR><BR> | ||
'''Service Status'''<BR> | '''Service Status'''<BR> | ||
<div style="background:#fff8e6; border-left:4px solid #c8960c; padding:8px 12px; margin-top:8px;">This service is in '''beta'''. The instrument is operational and BMC has completed initial runs, but experimental parameters are still being validated. We welcome early users willing to work collaboratively with BMC staff to establish protocols.</div> | <div style="background:#fff8e6; border-left:4px solid #c8960c; padding:8px 12px; margin-top:8px;">This service is in '''beta'''. The instrument is operational and BMC has completed initial runs, but experimental parameters are still being validated. We welcome early users willing to work collaboratively with BMC staff to establish protocols.</div> | ||
|} | |||
== SINGULAR G4X IN SITU SPATIAL SEQUENCING == | |||
<div style="background:#fff8e6; border-left:4px solid #c8960c; padding:8px 12px; margin-bottom:12px;">'''Beta Service — Early Access:''' The Singular G4X at the BioMicro Center is newly installed and not yet in routine service. '''Consultation with BMC staff is required before submitting samples.'''</div> | |||
<!-- placeholder image left: Singular G4X instrument photo or spatial multiomics schematic --> | |||
<!-- replace with: [[File:SingularG4X_instrument.png|thumb|left|400px]] --> | |||
The BioMicro Center provides access to the [https://www.singulargenomics.com/g4x Singular Genomics G4X], an upgrade to the G4 sequencer that adds in situ spatial multiomics capability. The G4X simultaneously profiles targeted RNA transcripts, proteins, and fluorescent H&E (fH&E) directly in intact FFPE tissue sections at subcellular resolution. Runs are organized around individual flow cells; BMC will typically run one flow cell at a time. Because this is an early-access service, results and run parameters are less well-established than our routine offerings. Consultation with BMC staff is strongly encouraged before designing experiments. | |||
<BR><BR> | |||
{| | |||
|- style="vertical-align: top;" | |||
|style="width: 500px;"| | |||
=== FLOW CELL OPTIONS (per slide) === | |||
{| class="wikitable" border=1 | |||
! Flow cell !! Lanes !! ROI size !! Samples !! Imaging area !! Run time | |||
|- | |||
! X2 | |||
| 2 || 10 × 10 mm || 10 || 10 cm² || ~29 hrs | |||
|- | |||
! X4 | |||
| 4 || 4.5 × 4.5 mm || 32 || 6.5 cm² || ~24 hrs | |||
|} | |||
<small>Run times do not include analysis time (~1 day per flow cell after imaging). Consult BMC staff to select the appropriate flow cell for your project.</small> | |||
=== PLATFORM SPECIFICATIONS === | |||
{| class="wikitable" border=1 | |||
! Instrument | |||
| [https://www.singulargenomics.com/g4x Singular Genomics G4X] | |||
|- | |||
! Spatial modalities | |||
| Targeted RNA (in situ), targeted protein, fluorescent H&E (fH&E) | |||
|- | |||
! RNA plex | |||
| Up to 500-plex per run | |||
|- | |||
! Protein plex | |||
| Up to 18-plex per run | |||
|- | |||
! Tissue format | |||
| FFPE (primary); other formats — consult BMC staff | |||
|- | |||
! Sample-to-answer | |||
| ~5 days | |||
|} | |||
For current pricing, see [[BioMicroCenter:Pricing]]. Details to be added as the service matures. | |||
=== SUBMISSION === | |||
<!-- iLabs link stub: replace once service item is created --> | |||
Submission via iLabs is not yet available. Contact BMC staff directly to discuss your experiment before scheduling. '''Consultation is required for all G4X submissions.''' | |||
| | |||
=== Singular G4X at BMC === | |||
<!-- placeholder image right: G4X example spatial data showing tissue section with RNA/protein overlay --> | |||
<!-- replace with: [[File:SingularG4X_example.png|thumb|right|350px]] --> | |||
The Singular G4X enables in situ spatial multiomics by co-detecting RNA transcripts, targeted proteins, and fluorescent H&E simultaneously in the same FFPE tissue section at subcellular resolution. Tissue sections are transferred to flow cells using Singular's TissuStamp™ system, and the instrument images the full flow cell in a single day. BMC will typically run one flow cell at a time, with flow cell choice (X2 or X4) determining sample capacity and ROI size. | |||
<BR><BR> | |||
The platform uses application-specific RNA panels (290–500 plex depending on tissue type) and a protein panel (up to 18-plex). Panels can be customized with add-ons of up to 50 genes, new 150-gene modules, or fully custom 500-gene panels. Current off-the-shelf panels include Lung, Colon, Breast, Kidney, Pancreas, Skin, and Pan-Mouse. See the [https://www.singulargenomics.com/content Singular content page] for the full panel list and gene targets. | |||
<BR><BR> | |||
For more detail on chemistry, workflow, and analysis, see the [https://www.singulargenomics.com/g4x-platform-ecosystem G4X Platform Ecosystem] and [https://www.singulargenomics.com/g4x G4X in situ multiomics] pages from Singular Genomics. | |||
<BR><BR> | |||
'''Service Status'''<BR> | |||
<div style="background:#fff8e6; border-left:4px solid #c8960c; padding:8px 12px; margin-top:8px;">This service is in '''beta'''. The G4X is newly installed and BMC is establishing protocols and validated run parameters. We welcome early users willing to work collaboratively with BMC staff. Contact biomicro@mit.edu to discuss your project.</div> | |||
|} | |} | ||
Latest revision as of 18:21, 17 April 2026
HOME -- SEQUENCING -- LIBRARY PREP -- HIGH-THROUGHPUT -- COMPUTING -- OTHER TECHNOLOGY
The BioMicro Center supports two complementary approaches to spatial transcriptomics: 10x Visium HD from 10x Genomics for slide-based spatial gene expression, and AVITI24 in situ sequencing from Element Biosciences. The choice of method depends on your tissue type, resolution requirements, and experimental goals. We strongly encourage consultation with BMC staff prior to beginning a spatial experiment.
10X VISIUM HD
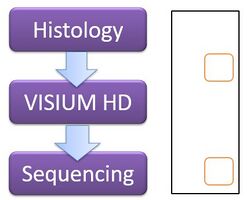 Workflow and capture areas for Visium HD
|
10x Visium HD is a spatial transcriptomics platform that profiles whole-transcriptome gene expression across intact tissue sections at single-cell-scale resolution. Tissue sections are processed through the CytAssist instrument, which transfers RNA complexes onto a capture slide containing a continuous array of 3,000,000 spatially barcoded 2 µm squares covering a 6.5 × 6.5 mm capture area (~0.65 cm²). Two chemistries are available: a probe-based assay for human and mouse (FF, FxF, and FFPE), and a polyA-based assay (Visium HD 3') for fresh frozen tissue from any species. H&E or immunofluorescence (IF) staining can be performed prior to processing for protein co-detection.
|
ARRAY SPECIFICATIONS
CHEMISTRY OPTIONS
TISSUE PREPARATION REQUIREMENTS
For current pricing, see BioMicroCenter:Pricing → Single Cell & Spatial Services → Visium. Sequencing is charged separately; 400–800M reads per tile recommended for Visium HD. SUBMISSIONSubmit via iLabs (assisted service). Coordinate with BMC staff at least one week in advance. Tissue processing begins at the Tang Histology Facility; please arrange sectioning separately. |
10X Visium HD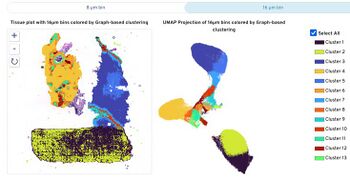 FAQs for Users | ||||||||||||||||||||||||||||||||||||||||||||
AVITI24 CYTOPROFILING & IN SITU SEQUENCING
The BioMicro Center provides access to the Element Biosciences AVITI24 platform for in situ single-cell multiomics. Using Teton™ CytoProfiling, the instrument co-detects RNA transcripts, proteins (including phospho-proteins), and cell morphology at subcellular resolution — directly in intact cells, without library preparation. Targeted in situ sequencing via Direct In Sample Sequencing (DISS) can be run simultaneously in the same experiment. Because this is an early-access service, results and run parameters are less well-established than our established offerings. Consultation with BMC staff is strongly encouraged before designing experiments.
PLATFORM SPECIFICATIONS
ASSAY CAPABILITIES
Teton CytoProfiling and DISS (low output) can be run simultaneously in the same experiment. Custom antibody add-ons require unconjugated rabbit monoclonal primaries. Consult BMC staff for current options. SAMPLE REQUIREMENTS
SUBMISSIONSubmission via iLabs is not yet available. Contact BMC staff directly to discuss your experiment before scheduling. Consultation is required for all AVITI24 Teton and DISS submissions. |
AVITI24 Teton CytoProfiling at BMC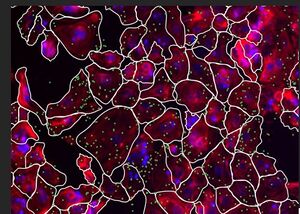 About the Service
This service is in beta. The instrument is operational and BMC has completed initial runs, but experimental parameters are still being validated. We welcome early users willing to work collaboratively with BMC staff to establish protocols.
| ||||||||||||||||||||||||||||||||||||||||
SINGULAR G4X IN SITU SPATIAL SEQUENCING
The BioMicro Center provides access to the Singular Genomics G4X, an upgrade to the G4 sequencer that adds in situ spatial multiomics capability. The G4X simultaneously profiles targeted RNA transcripts, proteins, and fluorescent H&E (fH&E) directly in intact FFPE tissue sections at subcellular resolution. Runs are organized around individual flow cells; BMC will typically run one flow cell at a time. Because this is an early-access service, results and run parameters are less well-established than our routine offerings. Consultation with BMC staff is strongly encouraged before designing experiments.
FLOW CELL OPTIONS (per slide)
Run times do not include analysis time (~1 day per flow cell after imaging). Consult BMC staff to select the appropriate flow cell for your project. PLATFORM SPECIFICATIONS
For current pricing, see BioMicroCenter:Pricing. Details to be added as the service matures. SUBMISSIONSubmission via iLabs is not yet available. Contact BMC staff directly to discuss your experiment before scheduling. Consultation is required for all G4X submissions. |
Singular G4X at BMCThe Singular G4X enables in situ spatial multiomics by co-detecting RNA transcripts, targeted proteins, and fluorescent H&E simultaneously in the same FFPE tissue section at subcellular resolution. Tissue sections are transferred to flow cells using Singular's TissuStamp™ system, and the instrument images the full flow cell in a single day. BMC will typically run one flow cell at a time, with flow cell choice (X2 or X4) determining sample capacity and ROI size.
This service is in beta. The G4X is newly installed and BMC is establishing protocols and validated run parameters. We welcome early users willing to work collaboratively with BMC staff. Contact biomicro@mit.edu to discuss your project.
|

