BioMicroCenter:SingleCell: Difference between revisions
Update AVITI24 hidden comment: high-output DISS not yet released |
|||
| (7 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
{{BioMicroCenter}} | {{BioMicroCenter}} | ||
The BioMicro Center supports | [[File:SingleCellOptions.png|thumb|right|400px]] | ||
The BioMicro Center supports a broad range of methods for single cell sequencing. The choice of method depends heavily on the type of question being asked and the source material. The methods break down into those supporting individual cells characterized in single wells and methods that use droplet isolation and library preparation. These cell requirements for each method vary significantly so reviewing each method is valuable and we strongly encourage consultation with BMC staff prior to beginning the experiment. | |||
<BR><BR><BR><BR><BR><BR>. | |||
== 10x CHROMIUM X == | == 10x CHROMIUM X == | ||
[[File:25_10xAssays.png|thumb|left|400px]] | |||
The BioMicro Center provides access to 10x Genomics library preparation as an assisted or a walk-up service. Added in 2023, The 10x Chromium X can handle a broad variety of methodologies now including [https://www.10xgenomics.com/solutions/single-cell/ 3', 5' and fixed RNA sequencing], [https://www.10xgenomics.com/solutions/single-cell-atac/ ATAC and multiome] <!-- [https://www.10xgenomics.com/solutions/single-cell-cnv/ CNV]. --> Dropoff for assisted service is coordinated with Center staff with at least one week's lead time. Users should bring their single cell/nuclei suspension(s) in 1.5 mL microfuge tubes in the standard buffer at or around that time. Staff works with the user to intake the initial samples and proceed through the protocol with quality control checks at the appropriate steps. | |||
<BR><BR> | |||
Chromium X usage is also offered as a walkup service. Usage may be scheduled on the iLabs calendar after training by BMC staff. This still requires at least a week's lead time before usage. You will not have permission to schedule the equipment until you have been approved by BMC staff. Reagents are stocked in the BioMicro Center and we ask that you use our reagents. This allows us to get larger bulk discounts we can pass on to our users. Please note that we do include a fraction of the instrument usage cost in the cost of the chips and will charge this cost if you use your own chips. | |||
<BR><BR> | |||
The full 10x suite of software is installed on LURIA and is integrated into our analysis package. <BR><BR> | |||
{| | {| | ||
|- style="vertical-align: top;" | |- style="vertical-align: top;" | ||
|style="width: 450px;"| | |style="width: 450px;"| | ||
=== | === scRNAseq / snRNAseq === | ||
<!-- A "setup" covers the chip, labor, and service contract. For Flex it also includes the GEMs. Samples per setup differ by assay - see Samples per setup row. --> | |||
{| class="wikitable" border=1 | {| class="wikitable" border=1 | ||
! | ! | ||
! 3'RNA/5'RNA <BR> (GEM-X) | ! 3'RNA/5'RNA <BR> (GEM-X) | ||
! 3'/5' OnChipMultiplexing <BR> (OCM) | ! 3'/5' OnChipMultiplexing <BR> (OCM) | ||
! Fixed RNA <BR> (Flex | ! Fixed RNA <BR> (Flex v2 / Apex) | ||
|- | |- | ||
! | !INPUT | ||
|colspan="2"| | |colspan="2"| | ||
[https://www.10xgenomics.com/support/universal-three-prime-gene-expression/documentation/steps/sample-prep Freshly counted cells/nuclei in suspension.] | [https://www.10xgenomics.com/support/universal-three-prime-gene-expression/documentation/steps/sample-prep Freshly counted cells/nuclei in suspension.] | ||
| | | | ||
[https://www.10xgenomics.com/support/flex-gene-expression/documentation/steps/sample-prep Fixed cells in suspension.] | [https://www.10xgenomics.com/support/flex-gene-expression/documentation/steps/sample-prep Fixed cells in suspension.] Also supports fresh, frozen, FFPE, fixed whole blood, and flow-sorted cells. | ||
|- | |||
!Cell Concentration <BR><small> cells|nuclei/uL </small> | |||
|100-2000 | |||
|100-2000 | |||
|100-2000 | |||
|- | |- | ||
! Fraction Recovered <BR><small> est. percent of input cells captured </small> | |||
| ~65% | |||
| ~60% | |||
| ~50% | |||
|- | |||
| | ! Target recovery <BR> <small> multiply by recovery for input </small> | ||
| 500-20,000/lane | |||
| 500-5,000/lane | |||
| up to 20,000/sample | |||
|- | |||
| | ! Expected doublet rate @ Max load | ||
| 8% | |||
| 7.5% | |||
| 8% doublet @ 20k | |||
|- | |- | ||
! | ! Samples per setup <BR><small> A setup covers chip, labor, and service contract. Flex also includes GEMs. </small> | ||
| 1 | | 1-8 samples <BR><small> 8 GEMs per chip </small> | ||
| 4 | | 1-8 samples <BR><small> 4 lanes/batch → 2 GEMs per chip </small> | ||
| | | up to ~48 samples <BR><small> 1 GEM; ~1M cells captured; ~20k cells/index </small> | ||
|- | |- | ||
!ADD'L AMPLICONS | !ADD'L AMPLICONS | ||
| Line 47: | Line 60: | ||
| | | | ||
Additional genes [https://www.10xgenomics.com/support/flex-gene-expression/documentation/steps/experimental-design-and-planning/custom-probe-design-for-visium-spatial-gene-expression-and-chromium-single-cell-gene-expression-flex require new probes.] | Additional genes [https://www.10xgenomics.com/support/flex-gene-expression/documentation/steps/experimental-design-and-planning/custom-probe-design-for-visium-spatial-gene-expression-and-chromium-single-cell-gene-expression-flex require new probes.] | ||
|- | |||
! KEY NOTES | |||
| | |||
| Batches of 4 samples | |||
| Human/Mouse only. <BR><small> Probe-based: exogenous sequences (GFP, reporters, etc.) not captured without custom probes. </small> | |||
|- | |- | ||
!SUBMISSION FORMS | !SUBMISSION FORMS | ||
| Line 57: | Line 75: | ||
|colspan="3"| [http://mit.edu/manoli/ Prof. Manolis Kellis] | |colspan="3"| [http://mit.edu/manoli/ Prof. Manolis Kellis] | ||
|} | |} | ||
=== CHROMATIN === | === CHROMATIN === | ||
{| class="wikitable" border=1 | {| class="wikitable" border=1 | ||
| Line 80: | Line 98: | ||
|colspan="2"| FASTQ, BAM, 10X QC, loupe file | |colspan="2"| FASTQ, BAM, 10X QC, loupe file | ||
|} | |} | ||
| | | | ||
=== 10X Genomics === | === 10X Genomics === | ||
[[image:10xX.jpg|thumb|right|500px|10x Chromium X]] | [[image:10xX.jpg|thumb|right|500px|10x Chromium X]] | ||
FAQs for Users <BR> | FAQs for Users <BR> | ||
'''1) What buffers in the final suspension are compatible with 10X applications?''' | '''1) What buffers in the final suspension are compatible with 10X applications?''' | ||
<br> | <br> | ||
[https://kb.10xgenomics.com/ | [https://kb.10xgenomics.com/s/article/115001937123-What-buffers-can-be-used-for-washing-and-cell-resuspension Buffers/media] for the submitted single cell/nuclei suspension should not contain excessive amounts of EDTA (>0.1 mM) or magnesium (>3 mM) and should be free of surfactants (i.e. Tween-20, SDS etc) and any RNases or DNases. | ||
*1xPBS (calcium free and magnesium free) containing 0.04% weight/volume BSA (400 µg/ml) is recommended for most general protocols and is considered the standard buffer | *1xPBS (calcium free and magnesium free) containing 0.04% weight/volume BSA (400 µg/ml) is recommended for most general protocols and is considered the standard buffer | ||
*Cell culture media with up to 1% BSA or up to 10% FBS if cells are not viable in standard buffer | *Cell culture media with up to 1% BSA or up to 10% FBS if cells are not viable in standard buffer | ||
| Line 100: | Line 114: | ||
*Nuclei also require addition of RNase Inhibitor along with 10X Genomics 1X nuclei buffer before chip loading, instructions for which are included in 10x user guides. If needed, users can collect buffer aliquots from the BMC after submitting a project. Please coordinate with BMC staff for pick-up. <br> | *Nuclei also require addition of RNase Inhibitor along with 10X Genomics 1X nuclei buffer before chip loading, instructions for which are included in 10x user guides. If needed, users can collect buffer aliquots from the BMC after submitting a project. Please coordinate with BMC staff for pick-up. <br> | ||
'''2) How many cells are captured in the Assay?''' | '''2) How many cells are captured in the Assay?''' | ||
<br>Up to | <br>Up to 20,000 cells per GEM can be uniquely barcoded with GEM-X, but this highly depends on cell counts and viability. Dying cells will leak RNA, hence may not be captured efficiently leading to sample failures. We recommend to count cells at the BMC to avoid discrepancies, but can work with users' counts as well. <BR> | ||
'''3) What are the best practices for flow sorting cells? <br> | '''3) What are the best practices for flow sorting cells?''' <br> | ||
10x provides guidance with their tested protocols about pre-sort buffer, collection buffer and FACS best practices [https://kb.10xgenomics.com/ | 10x provides guidance with their tested protocols about pre-sort buffer, collection buffer and FACS best practices [https://kb.10xgenomics.com/s/article/360048826911-What-are-the-best-practices-for-flow-sorting-cells-for-10x-Genomics-assays here.] <br> | ||
'''4) What is the expected size distribution for cDNA?''' <br> | '''4) What is the expected size distribution for cDNA?''' <br> | ||
cDNA for 3' and 5' libraries will span between 400 to 9000 base pairs, depending on sample type. | cDNA for 3' and 5' libraries will span between 400 to 9000 base pairs, depending on sample type. | ||
| Line 108: | Line 122: | ||
'''5) How much sequencing per sample is recommended?''' <br> | '''5) How much sequencing per sample is recommended?''' <br> | ||
10x makes several recommendations in their [https://www.10xgenomics.com/support/epi-atac/documentation/steps/sequencing/sequencing-handbook sequencing handbook]. Recommendation numbers vary by sample type, expected CNPs per sample, assay type and general sample quality. The higher the CNPs, the higher the quality, the more likely increased read depth is required. <br> | 10x makes several recommendations in their [https://www.10xgenomics.com/support/epi-atac/documentation/steps/sequencing/sequencing-handbook sequencing handbook]. Recommendation numbers vary by sample type, expected CNPs per sample, assay type and general sample quality. The higher the CNPs, the higher the quality, the more likely increased read depth is required. <br> | ||
For more resources, please visit https://kb.10xgenomics.com/ | For more resources, please visit https://kb.10xgenomics.com/s/ <br> | ||
| | | | ||
|} | |} | ||
== NAMOCELL SINGLE CELL SORTER == | == NAMOCELL SINGLE CELL SORTER == | ||
{| align=left | {| align=left | ||
|- style="vertical-align: top;" | |- style="vertical-align: top;" | ||
| Line 147: | Line 161: | ||
|} | |} | ||
| | | | ||
[[image:Namo.jpg|thumb|right|Namo (Namocell) Bio-Techne]] | [[image:Namo.jpg|thumb|right|Namo (Namocell) Bio-Techne]] | ||
The [https://www.bio-techne.com/brands/namocell Namocell Single Cell Sorter] allows users to sort cells into plates. The sorter uses microfluidics to sort single cells in 1uL of sheath fluid into a well. The instrument uses disposable cartridges to minimize contamination. The instrument integrates well with the [[BioMicroCenter:Tecan_Freedom_Evo#SPT_LABTECH_MOSQUITO_HV|SPT Labtech Mosquito HV]] which handles small reaction volumes. <BR><BR> | |||
|} | |} | ||
<BR><BR>. | |||
<!-- | |||
<!-- AVITI24 high-output DISS (whole transcriptome single cell) section pending - not yet released --> | |||
<!-- commenting out Seq-Well | <!-- commenting out Seq-Well | ||
== SEQ-WELL == | == SEQ-WELL == | ||
| Line 191: | Line 202: | ||
Seq-Well projects can be brought in two ways. For full service, please use the ilabs form linked on the left. At the end of the form it asks you to select several days on which the experiment can be done. The Nanowell Core will review the possible dates with you to fit it in the schedule. On that date, the core will confirm the quality of your cells prior to beginning the experiment. While 10,000 cells are typically used in a Seq-Well experiment, the core requests you plan to bring more cells - often counts from flow cytometers and other instruments are incorrect. Planning to bring more cells ensures you have the 10,000 desired. The BioMicro Center becomes involved in these projects at the exonuclease treatment step, where we take over the preparation of the library and quality control, as well as sequencing and analysis.<BR><BR> | Seq-Well projects can be brought in two ways. For full service, please use the ilabs form linked on the left. At the end of the form it asks you to select several days on which the experiment can be done. The Nanowell Core will review the possible dates with you to fit it in the schedule. On that date, the core will confirm the quality of your cells prior to beginning the experiment. While 10,000 cells are typically used in a Seq-Well experiment, the core requests you plan to bring more cells - often counts from flow cytometers and other instruments are incorrect. Planning to bring more cells ensures you have the 10,000 desired. The BioMicro Center becomes involved in these projects at the exonuclease treatment step, where we take over the preparation of the library and quality control, as well as sequencing and analysis.<BR><BR> | ||
Many laboratories are also using reagents from the Nanowell core to begin doing Seq-Well on their own. For these labs, we strongly recommend at least confirming the quality of the cDNA on our Advanced Analytical before continuing through library preparation. The library preparation can also be done in the BioMicro Center. Please use the standard Illumina library preparation form and note that the samples are Seq-Well. We do have all the custom primers needed for preparation and the pricing is identical to other NexteraXT preps. | Many laboratories are also using reagents from the Nanowell core to begin doing Seq-Well on their own. For these labs, we strongly recommend at least confirming the quality of the cDNA on our Advanced Analytical before continuing through library preparation. The library preparation can also be done in the BioMicro Center. Please use the standard Illumina library preparation form and note that the samples are Seq-Well. We do have all the custom primers needed for preparation and the pricing is identical to other NexteraXT preps. | ||
|} | |} | ||
--> | --> | ||
Latest revision as of 20:29, 24 March 2026
HOME -- SEQUENCING -- LIBRARY PREP -- HIGH-THROUGHPUT -- COMPUTING -- OTHER TECHNOLOGY
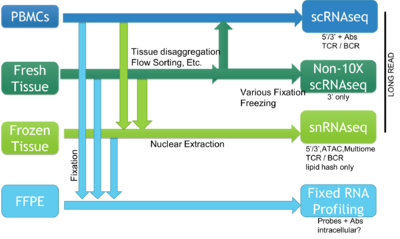
The BioMicro Center supports a broad range of methods for single cell sequencing. The choice of method depends heavily on the type of question being asked and the source material. The methods break down into those supporting individual cells characterized in single wells and methods that use droplet isolation and library preparation. These cell requirements for each method vary significantly so reviewing each method is valuable and we strongly encourage consultation with BMC staff prior to beginning the experiment.
.
10x CHROMIUM X

The BioMicro Center provides access to 10x Genomics library preparation as an assisted or a walk-up service. Added in 2023, The 10x Chromium X can handle a broad variety of methodologies now including 3', 5' and fixed RNA sequencing, ATAC and multiome Dropoff for assisted service is coordinated with Center staff with at least one week's lead time. Users should bring their single cell/nuclei suspension(s) in 1.5 mL microfuge tubes in the standard buffer at or around that time. Staff works with the user to intake the initial samples and proceed through the protocol with quality control checks at the appropriate steps.
Chromium X usage is also offered as a walkup service. Usage may be scheduled on the iLabs calendar after training by BMC staff. This still requires at least a week's lead time before usage. You will not have permission to schedule the equipment until you have been approved by BMC staff. Reagents are stocked in the BioMicro Center and we ask that you use our reagents. This allows us to get larger bulk discounts we can pass on to our users. Please note that we do include a fraction of the instrument usage cost in the cost of the chips and will charge this cost if you use your own chips.
The full 10x suite of software is installed on LURIA and is integrated into our analysis package.
scRNAseq / snRNAseq
CHROMATIN
|
10X Genomics
2) How many cells are captured in the Assay?
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
NAMOCELL SINGLE CELL SORTER
|
 The Namocell Single Cell Sorter allows users to sort cells into plates. The sorter uses microfluidics to sort single cells in 1uL of sheath fluid into a well. The instrument uses disposable cartridges to minimize contamination. The instrument integrates well with the SPT Labtech Mosquito HV which handles small reaction volumes. |
.

